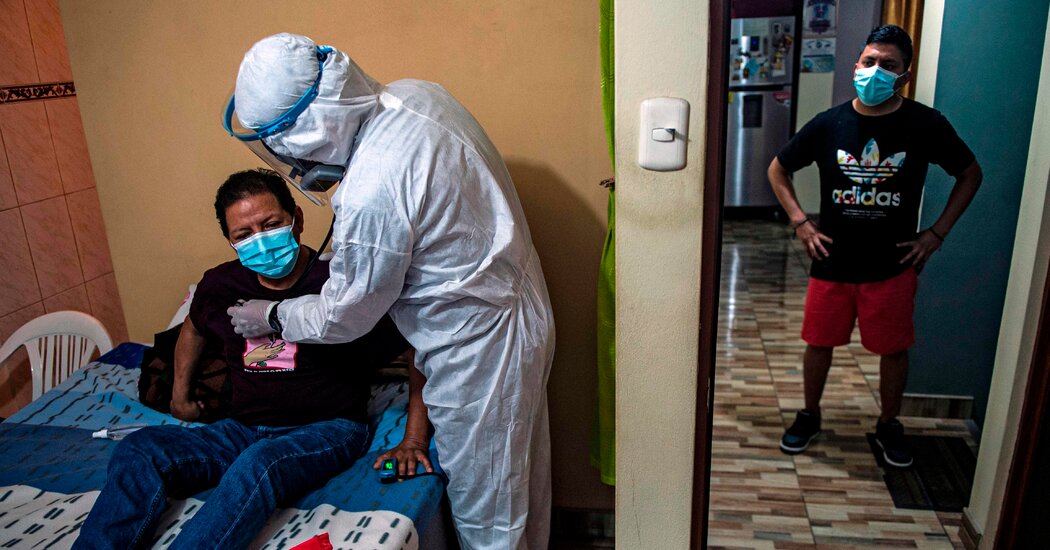
Spreading fast in South America, the variant is still a mystery. No one knows whether it is more contagious than other variants or if it affects vaccines.Viruses evolve. SARS-CoV-2, the virus that causes Covid-19, is no exception. So the emergence of variants is no surprise, and not every new genetic mutation poses a serious threat.But in recent weeks, a growing drumbeat of news coverage has started to raise alarm about Lambda, a variant first detected in Peru late last year. The variant, initially known as C.37, has spread rapidly through parts of South America. On June 14, the World Health Organization designated it as a “variant of interest,” meaning, essentially, that experts suspect it could be more dangerous than the original strain.Lambda’s prevalence and its mutations, which resemble those found in several other highly contagious or worrisome variants, mean that it is worth watching, scientists said. But much remains unknown, and it is not yet clear how much of a risk it poses.“I think some of the interest is just based on the fact that there’s a new variant, and it has a new name,” said Nathaniel Landau, a microbiologist at the New York University Grossman School of Medicine who is studying the new coronavirus variants.“But I don’t think there’s any more reason to be concerned than before we knew about this variant,” Dr. Landau added. No evidence so far suggests that Lambda will outcompete Delta, the highly transmissible variant that’s now dominating most of the world. “There’s no reason to think that this is now something worse than Delta.”Pablo Tsukayama, a microbiologist at Cayetano Heredia University in Peru who documented Lambda’s emergence, concurred. Latin America has “limited capacity” to do genomic surveillance and follow-up laboratory investigations of new variants, he said. That has led to an information gap fueling concerns about Lambda. “I don’t think it’s going to be worse than any of the ones that we have already,” he said. “It’s just that we know so little that it lends itself to a lot of speculation.”As of mid-June, Lambda had been reported in 29 countries, territories or areas, according to a June 15 update from the W.H.O. The variant had been detected in 81 percent of coronavirus samples sequenced in Peru since April, and 31 percent of those in Chile to date, the agency said.The variant accounts for less than 1 percent of samples sequenced in the United States, according to GISAID, a repository for viral genome data. Isolated cases have been reported in a number of other countries.The variant contains eight notable mutations, including seven in the gene for the spike protein, found on the surface of the virus. Some of these mutations are present in other variants and might make the virus more infectious or help it evade the body’s immune response.But big questions remain unanswered. It is not yet clear whether Lambda is more transmissible than other variants, whether it causes more severe disease or whether it renders vaccines less effective.“We don’t have a lot of information, compared to the other variants,” said Ricardo Soto-Rifo, a virologist at the University of Chile who has studied Lambda.Preliminary laboratory studies, which have not yet been published in peer-reviewed journals, provide reason for both concern and reassurance. In these studies, research teams led by Dr. Soto-Rifo and Dr. Landau found that antibodies induced by the Pfizer, Moderna and CoronaVac vaccines are less powerful against Lambda than against the original strain, but that they are still able to neutralize the virus.The findings suggest that these vaccines should still work against Lambda, the scientists said. Moreover, antibodies are not the body’s only defense against the virus; even if they’re less potent against Lambda, other components of the immune system, like T cells, may also provide protection.“This decrease in the neutralizing antibodies does not mean that the vaccine has decreased effectiveness,” Dr. Soto-Rifo said. Real-world studies of how well the vaccines hold up against the variant are still needed, he said.The researchers also reported that like several other variants, Lambda binds more tightly to cells than the original strain of the virus does, which may make it more transmissible.Although many questions remain, Trevor Bedford, an evolutionary biologist at the Fred Hutchinson Cancer Research Center in Seattle, said that he does not find Lambda as worrisome as Delta and does not expect it to become as dominant globally.“Lambda has been around for a little while, and it’s hardly invaded the U.S. at all, for example, compared to, say, even Gamma” — the variant first identified in Brazil — “which has done pretty well here.” He added, “I think all the focus should be on Delta.”
Read more →